Data were analyzed by means of unpaired t tests between groups and one-way analysis of variance to compare treatments within groups, with significant interactions further analyzed with the use of Tukey post hoc analysis with Bonferroni correction. (A) Levels of MPO mRNA were determined with the use of real-time reverse-transcription polymerase chain reaction, and (B) levels of MPO protein were detected with the use of ELISA. Human myometrial and leiomyoma cells were treated with and without dichloroacetate (DCA) (20 μg/mL) and with and without hypoxia (2% O 2, 5% CO 2-nitrogen gas balance) for 24 hours. A specific standard for each gene allows for absolute quantification of the gene in number of copies, which can then be expressed per microgram of RNA.įigure 1 Levels of myeloperoxidase (MPO) in myometrial and leiomyoma cells.

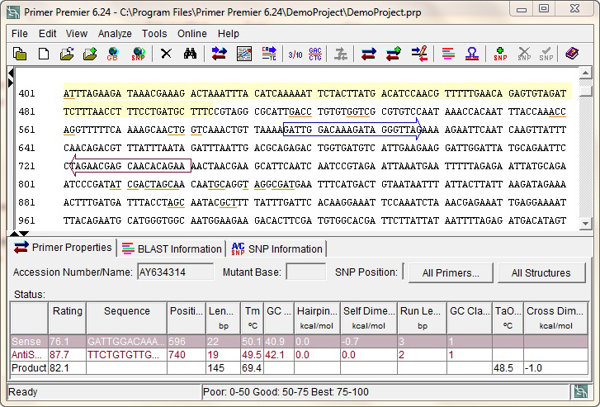
Standards with known concentrations and lengths (base pairs ) were designed specifically for β-actin (79 bp), Bax (85 bp), Bcl-2 (75 bp), MPO (106 bp), and iNOS (103 bp) with the use of the Beacon Designer software, allowing for construction of a standard curve using a tenfold dilution series. Real-time RT-PCR was performed in a 25-μL total reaction volume including 12.5 μL Express Sybr GreenER qPCR Supermix, 1 μL cDNA template, and 0.2 μmol/L each of target-specific primers designed to amplify a part of each gene. Sequences of the oligonucleotides used for amplification of β-actin, Bax, Bcl-2, MPO, and iNOS mRNA are described in Supplemental Table 1 (available online at Quantitative RT-PCR was performed with the use of the Express Sybr GreenER qPCR Supermix Kit (Life Technologies) and the Cepheid 1.2f detection system.
PREMIER BIOSOFT BEACON DESIGNER 8.1 SOFTWARE
Optimal oligonucleotide primer pairs for real-time reverse-transcription polymerase chain reaction (RT-PCR) amplification of reverse-transcribed cDNA were selected with the aid of the software program Beacon Designer (Premier Biosoft).
